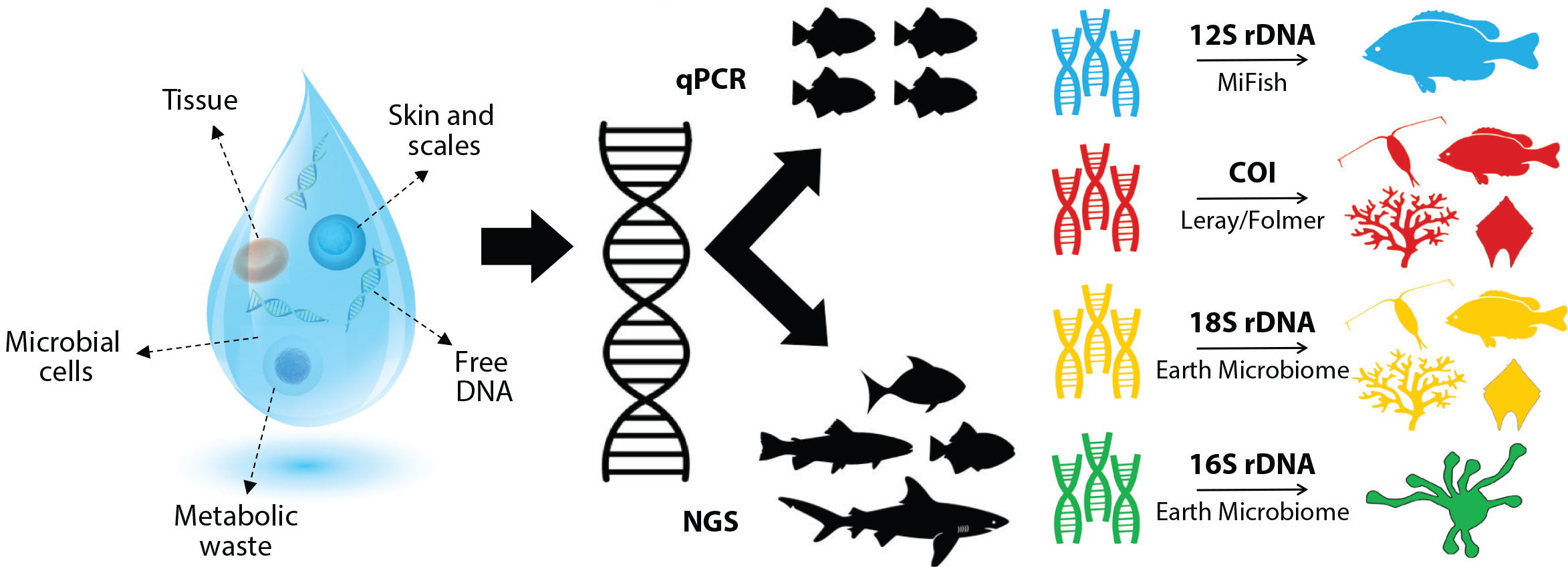
The scooping pond water and instantly knowing every fish, frog, insect, algae, and microbe living there. Metabarcoding makes this possible by reading environmental DNA (eDNA) from a single sample.It includes the following steps, Collect water, soil, or air. Extract all DNA mixed together. Then amplify specific barcode genes COI for animals, 16S for bacteria, ITS for fungi. Sequence millions of fragments. Match them to databases. It investigates that hundreds of species identified in days, with relative abundances.
Traditional surveys use nets and traps to catch 20-50 obvious species. Metabarcoding finds 300+ including microscopic life and rare individuals missed by sampling. It’s weatherproof, needs no expert taxonomists, and detects extracellular DNA from dead organisms. Stream monitoring shows its power. German studies found metabarcoding identified 3x more taxa than kick nets, including pollution-sensitive larvae. Forest soil samples reveal 5,000+ fungi controlling nutrient cycles. Invasive species like Asian clams get detected weeks before visual sightings. The real insight comes from traits. High shredder insect DNA predicts fast leaf breakdown rates. Fungal to bacterial ratios forecast carbon storage. Metabarcoding scales individual traits to entire ecosystem functions.
Challenges remain: 30% sequences match unknown dark taxa. PCR amplification biases favor certain species. Solutions include multi marker approaches and mock community calibration. Long term research sites like Hubbard Brook and Konza Prairie now integrate metabarcoding with 40 year baselines, tracking biodiversity shifts against climate and pollution trends. One sample reveals the full cast of ecosystem engineers, visible and invisible driving processes policymakers need to understand.